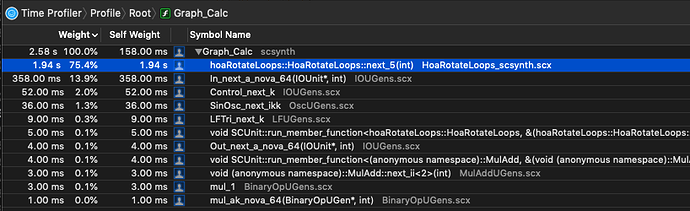
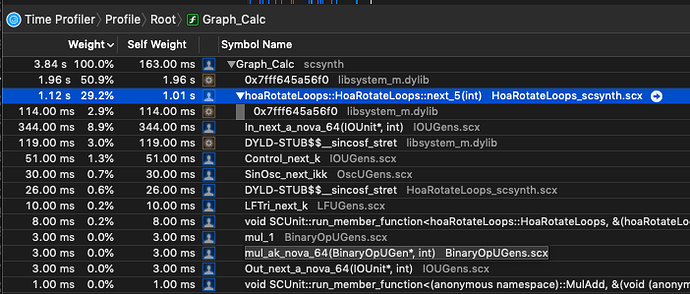
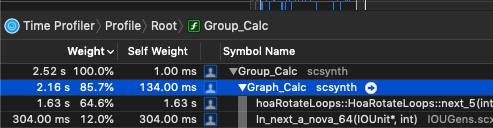
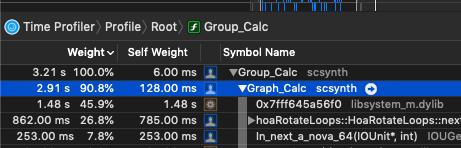
As I am beginning to write UGen’s, I am also trying to implement SIMD versions to get the most out them.
Two questions have come up:
- It seems that in my case
sin()andcos()are significantly slower on nova vectors—vec<float>— vs. simply iterating through each sample and calculating sin/cos on each sample. (MacBookPro, 2.9 GHz Intel Core i7)
const int vs = nova::vec<float>::size;
const int loops = nSamples / vs;
for (int i = 0; i < loops; ++i)
{
vec<float> r, r2, sinr, cosr, cosrm2, sinr2, cosr2;
r.load_aligned(rotation);
r2 = r * 2;
sinr = sin(r);
cosr = cos(r);
cosrm2 = cosr * 2;
sinr2 = sin(r2);
cosr2 = cos(r2);
cosrm2.store_aligned(cm2);
sinr.store_aligned(s);
cosr.store_aligned(c);
sinr2.store_aligned(s2);
cosr2.store_aligned(c2);
rotation += vs;
cm2 += vs;
s += vs;
c += vs;
s2 += vs;
c2 += vs;
}
(rotation, cm2, s, etc. are float * for data buffers, but could be I/O buffers, etc…)
While I haven’t meticulously isolated the cost of sin/cos on these vectors, in the context of the UGen I’m writing, simply swapping out a basic sample-iterating pattern that doesn’t use nova::vec:
for (int frm = 0; frm != nSamples; ++frm)
{
float r = *rotation++;
float r2 = r * 2;
float cosr = cos(r);
*s++ = sin(r);
*c++ = cosr;
*cm2++ = cosr * 2;
*s2++ = sin(r2);
*c2++ = cos(r2);
}
shows that it’s twice as fast. This seems counterintuitive, as I typically see a pretty decent speedup on things like SIMD binary operations.
Is this expected? How this is implemented is a bit obscure to me…
Is there something obviously wrong with how I’m using nova::vec?
I would imagine specific performance is architecture dependent.
If there is indeed a slowdown, this might be affecting other operations?
- This is a speculative question about whether this exists or could be implemented (feature request): Is there something like a “SIMD pointer type” for which there could be defined a custom iterator which steps
nova::vec<float>::size?
Such an iterator could make code like that above more concise:
const int vs = nova::vec<float>::size;
const int loops = nSamples / vs;
for (int i = 0; i < loops; ++i)
{
vec<float> r, r2, sinr, cosr, cosrm2, sinr2, cosr2;
r.load_aligned(rotation++);
r2 = r * 2;
sinr = sin(r);
cosr = cos(r);
cosrm2 = cosr * 2;
sinr2 = sin(r2);
cosr2 = cos(r2);
cosrm2.store_aligned(cm2++);
sinr.store_aligned(s++);
cosr.store_aligned(c++);
sinr2.store_aligned(s2++);
cosr2.store_aligned(c2++);
}
… and potentially get a performance boost?
Thanks for any insights!